 
You can find data in Pfam in various ways. In order to align sequences in SnapGene you should open your sequence and then select Tools-Align Multiple Sequences in the main menu (Figure 3.4.10.1). Overlapping is an alignment problem, but different from those we have discussed. Xgene & proteinX 263 subscribers Subscribe Share Save 1K views 1 year ago Align sequencing result Offline using snapgene ncbi. all UniProt and NCBI GI) or different levels of redundancy. YouTube 0:00 / 7:52 Align sequencing result using snapgene. Pfam full alignments are available from searching a variety of databases, either to provide different accessions (e.g. 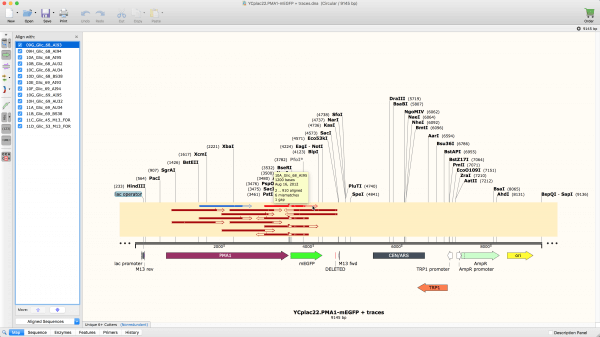
It contains minimal data and enables us to work easily with the alignment. Step 2 Choose any one family having less number of seed value. It will show all the Pfam families in alphabetical order. The data presented for each entry is based on the UniProt Reference Proteomes but information on individual UniProtKB sequences can still be found by entering the protein accession. Step 1 Open your favorite browser and go to website. 1) Get your sequences in FASTA format: Go to the NCBI website. Perfect conservation yields a value of 100. (Thomas Weimbs, University of California Santa Barbara, 11/2012). A clan is a collection of Pfam entries which are related by similarity of sequence, structure or profile-HMM. Alignment and Assembly How does SnapGene calculate conservation scores for a protein multiple alignment 8 months ago A conservation score for each position in a protein multiple alignment is calculated by the method of Valdar 1,2. Pfam also generates higher-level groupings of related entries, known as clans. The live, interactive views in SeqBuilder Pro make it effortless to edit and annotate your sequences, customize your features and create plasmid maps. The identification of domains that occur within proteins can therefore provide insights into their function. Different combinations of domains give rise to the diverse range of proteins found in nature. This tool is designed primarily for assembly of a small set of Sanger reads, all derived from the same clonal source, and all of which are expected to overlap to form a contiguous sequence. Proteins are generally composed of one or more functional regions, commonly termed domains. The SnapGene 'Assemble Contigs' tool uses the CAP3 assembler to assemble reads into one or more contiguous assemblies. The Pfam database is a large collection of protein families, each represented by multiple sequence alignments and hidden Markov models (HMMs). 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
